Originally developed to visualise complex neural circuits, Brainbow fluorescent protein (XFP) antibodies have become a powerful tool for tracking individual cells, amplifying multicolour signals, and enabling discoveries across tissues and organisms. We spoke with Prof. Dawen Cai from the University of Michigan, the inventor of these polyclonal antibodies against XFPs originated from very distinct spieces, to hear why he developed them and the impact they are having in both neuroscience and cancer research. By making these antibodies widely available, Prof. Cai is helping researchers to build on proven innovation, focus their efforts on new discoveries and accelerating progress across multiple fields.

Dawen Cai, PhD. University of Michigan
Seeing the brain clearly
Neuroscience is rapidly advancing through new antibodies and imaging technologies, allowing researchers to visualise the brain in remarkable detail. These insights are reshaping our understanding of how the brain develops, functions, and fails in disease.
For the cancer research community, this progress is increasingly relevant. Tumours often interact with surrounding tissue, including neural networks in the brain and peripheral nervous system. For example, glioblastoma research has shown that tumours do not invade the brain randomly and instead, hijacks the brain’s own neuronal mechanisms through malignant crosstalk, transforming normal brain function into an ecosystem for tumour progression (1). In brain cancers and in cancers that invade or communicate with nerves, understanding how malignant cells interact with neurons and their microenvironment requires imaging tools that can resolve neuronal structure and functions within living tissue.
Despite advances, a fundamental technical challenge persists: how can we visually distinguish individual neurons and cancer cells when they are packed together in dense, anatomically complex tissue? This limitation affects both basic neuroscience and translational cancer biology, where mapping cellular interactions at scale is essential.
The limits of traditional neural labelling
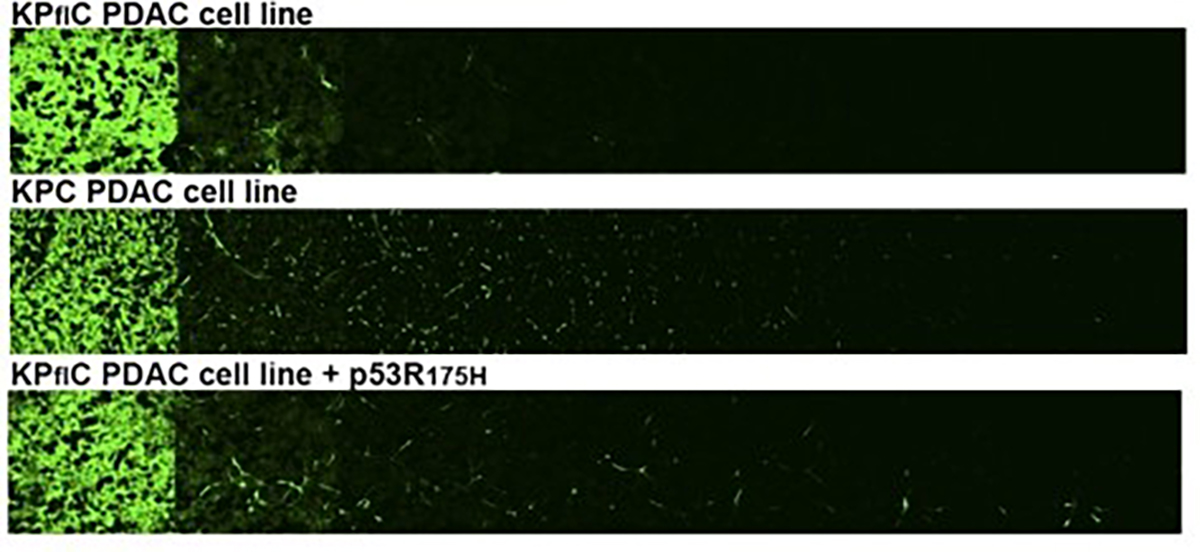
Traditional labelling methods, such as Golgi staining or dye injection, can reveal the morphology of a handful of neurons at a time, but never provide the complete picture. Fluorescent protein approaches have expanded what researchers can see, but still often result in weak signals, poor labelling of fine structures, and limited colour diversity. As a result, many studies have been forced to choose between imaging a small number of neurons clearly or many neurons indistinctly — a trade-off that has slowed progress in understanding both brain circuitry and disease, including cancer-related changes in the nervous system.
A major advancement came in 2007 with the introduction of Brainbow technology, developed by Jeff Lichtman and Josh Sanes at Harvard. By engineering transgenic mice to express different combinations of fluorescent proteins, researchers could label individual neurons in ~ 100 distinct colour combinations. This transformed the study of neural connectivity and made large-scale neural circuit mapping possible (2).
However, during Prof. Cai’s time at Harvard, he and his colleagues became aware of practical limitations that were preventing Brainbow from reaching its full potential. In particular:
- The original fluorescent proteins were prone to photoinstability and often produced weak or inconsistent signals.
- Because the proteins were cytoplasmic, they accumulated in neuronal cell bodies, failing to adequately label delicate axons and dendrites where critical connectivity information resides.
- The non-recombinant state expressed red fluorescent protein (RFP), producing a dominant “default” red colour that reduces true diversity across labelled neurons.
- The fluorescent proteins shared too much sequence homology to allow the design of specific immunostaining antibodies, making it impossible to recover signal once native fluorescence was lost during fixation, sectioning or tissue clearing.
To overcome these limitations, Prof. Cai and his team developed the Brainbow XFP antibodies, creating a new generation of tools that make multicolour neural imaging brighter, more reliable, and more broadly applicable.
”The Brainbow antibodies solve a fundamental bottleneck in multicolor fluorescent labeling: they provide a robust, species-agnostic way to amplify and recover the fluorescent signal after tissue fixation or processing, which is essential for any multicolor labeling experiment to be reliably quantified and compared across labs.
Dawen Cai, PhD

Cholenergic neurons labelled by AAV-Brainbow and stained with Brainbow XFP antibodies against mTFP, mCherry, TagRFP, and EGFP. Image credit: Dr. Douglas Roossien.
From Brainbow 1 to Brainbow 3: designing better fluorophores
Prof. Cai developed Brainbow 3 with the selection of improved XFPs that were brighter, more photostable, and less prone to aggregation than those used in earlier versions. As a result, the original fluorophores were replaced with mOrange2 (from coral), EGFP (from jellyfish), and mKate2 (from sea anemone), significantly improving signal quality and stability.
The team also engineered farnesylated derivatives of these proteins, so they localised to the cell membrane rather than accumulating in the cell body. This adaptation dramatically improved labelling of fine neuronal structures, including axons and dendrites — features critical for mapping neural connectivity.
Building on this, Prof. Cai’s group expanded the colour palette further through new transgene designs, Brainbow 3.1, 3.2, and AAV-Brainbow. By incorporating a broader range of XFPs, the system supported more distinct colours, eliminated the dominant “default red” problem, and enabled clearer discrimination between neighbouring neurons. Regulatory elements, including “stop” cassettes, were also introduced to prevent default expression of a single fluorophore and promote a more balanced colour distribution across labelled cells.
The key challenge: achieving true antibody specificity
Developing Brainbow 3 was not straightforward. Fluorescent proteins belong to a relatively conserved family, meaning that antibodies raised against closely related proteins often cross-react – a problem that would undermine multicolour labelling strategy. For immunostaining to work alongside Brainbow, each antibody needed to cleanly distinguish one fluorophore from all others.
The solution was strategic fluorescent protein selection. By choosing proteins from diverse evolutionary lineages – including coral, jellyfish, and sea anemones – the team maximised sequence divergence and antigenic distance. This enabled the generation of highly specific antibodies with minimal cross-reactivity. Ultimately, the panel comprised eight spectrally and antigenic distinct fluorescent proteins: mTFP1, EGFP, mNeonGreen, TagYFP, PhiYFP, mKusabiraOrange2, TagRFP, and mCherry.
Making the antibodies work in practice
A further challenge was ensuring the antibodies performed reliably across real-world experimental settings. Brainbow is used in transgenic mice, via AAV vectors, in zebrafish and Drosophila, and across a wide range of tissue processing workflows. To maximise compatibility, Prof. Cai’s team generated <strong>custom polyclonal antibodies in multiple host species — including chicken, rabbit, rat, and guinea pig — giving researchers more flexibility in choosing a broad range of secondary antibodies.
These antibodies were validated in cells and mice to confirm that they amplified signals without compromising colour diversity. Crucially, the team demonstrated that antibody amplification enhanced signal strength while preserving the spectral distinctions required for multicolour analysis — a result that underpinned their broad adoption (3).

Transgenic Bitbow Drosophila larval brain. The novel Bitbow design integrates five antigenic distinct FPs (mAmetrine, mTFP1, mNeonGreen, mKusabiraOrange2, and tdKatushka2) that are targeted to the nucleus, Golgi aparatus, and cytoplasmic membrane. After Flp induced random recombination, the 15-bit binary subcellular spectral barcodes are stained with the Brainbow XFP antibodies and utilised for neuron lineage tracing. Image credit: Dr. Ye Li
The outcome: a robust, versatile Brainbow XFP antibody toolkit
This improved antibody panel became the critical enabling technology. It transformed the original Brainbow from a technique reliant solely on native fluorescence into a robust, versatile immunostaining toolkit that performs reliably across fixation, tissue clearing, expansion microscopy, and a wide range of experimental settings.
”For any researcher working with fluorescent protein-based cell labelling — whether in neuroscience, cancer biology, or developmental biology — the Brainbow antibodies transform a technically fragile approach into a dependable, standardised toolkit.
Dawen Cai, PhD
From one lab to the world: sharing the Brainbow for faster discovery
Accessing research tools developed in other academic labs has normally involved lengthy email exchanges, complex material transfer agreements, and unpredictable shipping timelines — all of which can slow scientific progress. Through the non-profit research tool platform CancerTools, access to <strong>Brainbow XFP antibodies is now streamlined and reliable, reducing these practical barriers and enabling researchers to adopt validated tools more quickly.
As Prof. Cai explained, the Brainbow XFP antibodies were initially developed to solve a technical challenge within his own laboratory, but it soon became clear that this problem was universal. By making these antibodies available through CancerTools, laboratories worldwide can access a proven, well-characterised solution without having to recreate and optimise from scratch, an approach that reflects our shared mission with Cancer Research UK.
By enabling wide access to Brainbow XFP antibodies, Prof. Cai has helped cultivate a culture of openness and collaboration, ensuring that his work continues to empower researchers across neuroscience and cancer biology.
”Personally, it has reinforced my belief that the most impactful thing a methodologist can do is not just develop tools but make them accessible. The barrier to adoption matters enormously. A brilliant tool that sits in one lab is worth far less than a good tool that hundreds of labs can pick up and use.
Dawen Cai, PhD
From brain circuits to cancer discovery
In neuroscience, Brainbow antibodies continue to support high-resolution circuit mapping by preserving signal through fixation, tissue clearing, and advanced imaging — capabilities that now extend beyond the brain.
What has been particularly striking is how the Brainbow framework has inspired new approaches in cancer biology. A leading example is the Cancer Rainbow (Crainbow) mouse model developed by the Snyder laboratory at Duke University. Building directly on Brainbow’s principle of combinatorial fluorescent barcoding, the team paired specific oncogenic mutations, such as β-catenin (Ctnnb1), with distinct fluorescent proteins, allowing visualisation of clonal expansion in living tissue (4). Their work revealed how developmental timing and microenvironmental signals such as oncogenic RSPO3 can drive the emergence and spread of premalignant clones in adult intestines, helping explain tumour heterogeneity and early cancer evolution.
While Brainbow antibodies were not the primary tool used in Crainbow, the multicolour fluorescent protein principles pioneered by Brainbow made experiments like this possible. Equally, antibody-based signal amplification remains essential whenever robust detection is required in fixed or processed tissues. This is a powerful example of how one foundational technology can seed entirely new innovations in cancer research.
”It has been deeply rewarding to see the Brainbow antibodies adopted by labs working on questions I would never have imagined when we first developed them. The moment that really crystallized this for me was learning about the Crainbow work at Duke — seeing how the principles of multicolor fluorescent barcoding had been adapted to directly visualise the spread of oncogenic mutations in the gut, and how that led to the surprising finding that colorectal cancer may be seeded during early postnatal development. That is the kind of impact that makes tool development feel truly worthwhile — when a technology you built opens a door into a completely new area of biology.
Dawen Cai, PhD
Brainbow in the years ahead
Brainbow antibodies are more than a neuroscience tool – they form a general-purpose toolkit for experiments that rely on stochastic multicolour fluorescent labelling. For cancer researchers, they represent a powerful opportunity to visualise clonal evolution, tumour heterogeneity, and interactions with the tumour microenvironment, opening new approaches to study how cancers arise, diversify, and progress at the cellular level.
The development and widespread adoption of Brainbow 3 by Prof. Dawen Cai exemplify how thoughtfully designed research tools can shape the future of science. By advancing neuroscience while enabling innovation in cancer biology, his work provides a foundation that researchers can build on — supporting new insights and discoveries for years to come.

Parvalbumin expressing neurons labelled by AAV-Brainbow and stained with Brainbow 3 XFP antibodies against mTFP, mCherry, TagRFP, and EGFP. Image credit: Dawen Cai
Explore the Brainbow antibodies to power your next discovery
References:
- Venkataramani, V., et al. 2022. Cell. 185(16), 2899-2917. PMID: 35914528
- Livet, J., et al. 2007. Nature. 450, 56-62. PMID: 17972876
- Cai, D., et al. 2013. Nature Methods. 10, 540-547. PMID: 23817127
- Boone, P.G., et al. 2019. Nature Communications. 10, 5490.PMID: 31792216