St George’s University of London and the Functional Genomics Cell Bank strengthen their partnership with CancerTools.org to accelerate cancer research.
St George’s University of London (SGUL) and CancerTools.org announce new cell lines from Dr. Elena Sviderskaya, which are now readily available via the CancerTools.org website.
CancerTools.org is a global, non-profit cancer-focused biorepository with a 40+ year history of making cancer research tools accessible such as antibodies, cell lines, mouse models and more to cancer researchers worldwide.
Leading cancer biologists at SGUL are supporting CancerTool.org’s mission to accelerate cancer research through research tools. Since December 2017, CancerTools.org has been storing, producing and shipping SGUL’s scientists’ research tools worldwide. To accelerate cancer research and discoveries globally, SGUL and the Functional Genomics Cell Bank are now making 68 murine melanocyte and melanoblast cell lines, invented by Dr. Elena Sviderskaya, available through CancerTools.org.
“As a non-profit, CancerTools.org is dedicated to accelerating cancer breakthroughs, by ensuring cancer scientists have access to the highest quality research tools. We are delighted to be able to make these valuable cell lines available to the global research community and to continue strengthening our relationship with SGUL and the Functional Genomics Cell Bank.”
James Ritchie, Head of External Innovation, CancerTools.org
Dr. Elena Sviderskaya specialises in pigment cell and melanoma research1 and is the Director of the Functional Genomics Cell Bank1 at St George’s, University of London. The Functional Genomics Cell Bank specialises in mouse melanocyte and melanoblast lines carrying a variety of pigmentary mutations, immortal human melanocytes, melanoma cell lines, and stem cells. Many of these lines are now available through Cancer.Tools.org.
“We are delighted that the cell lines held in the Functional Genomics Cell Bank at St George's are now available via CancerTools.org to researchers globally. These lines are important for cancer biologists as non-cancerous controls for the behaviour of melanoma lines, and are also useful in testing the importance of melanogenesis in the progression of melanoma and other skin cancers.”
Dr. Elena Sviderskaya, Director of the Functional Genomics Cell Bank
About the cell lines:
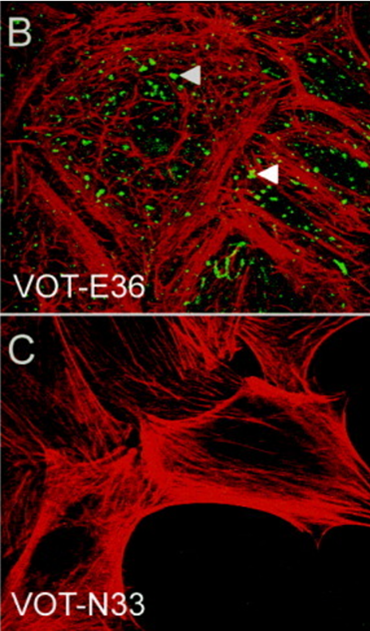
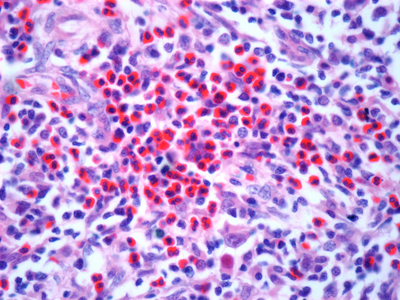
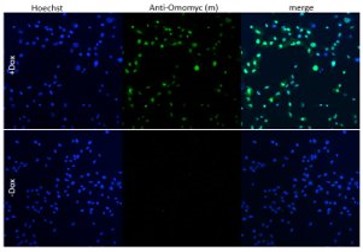
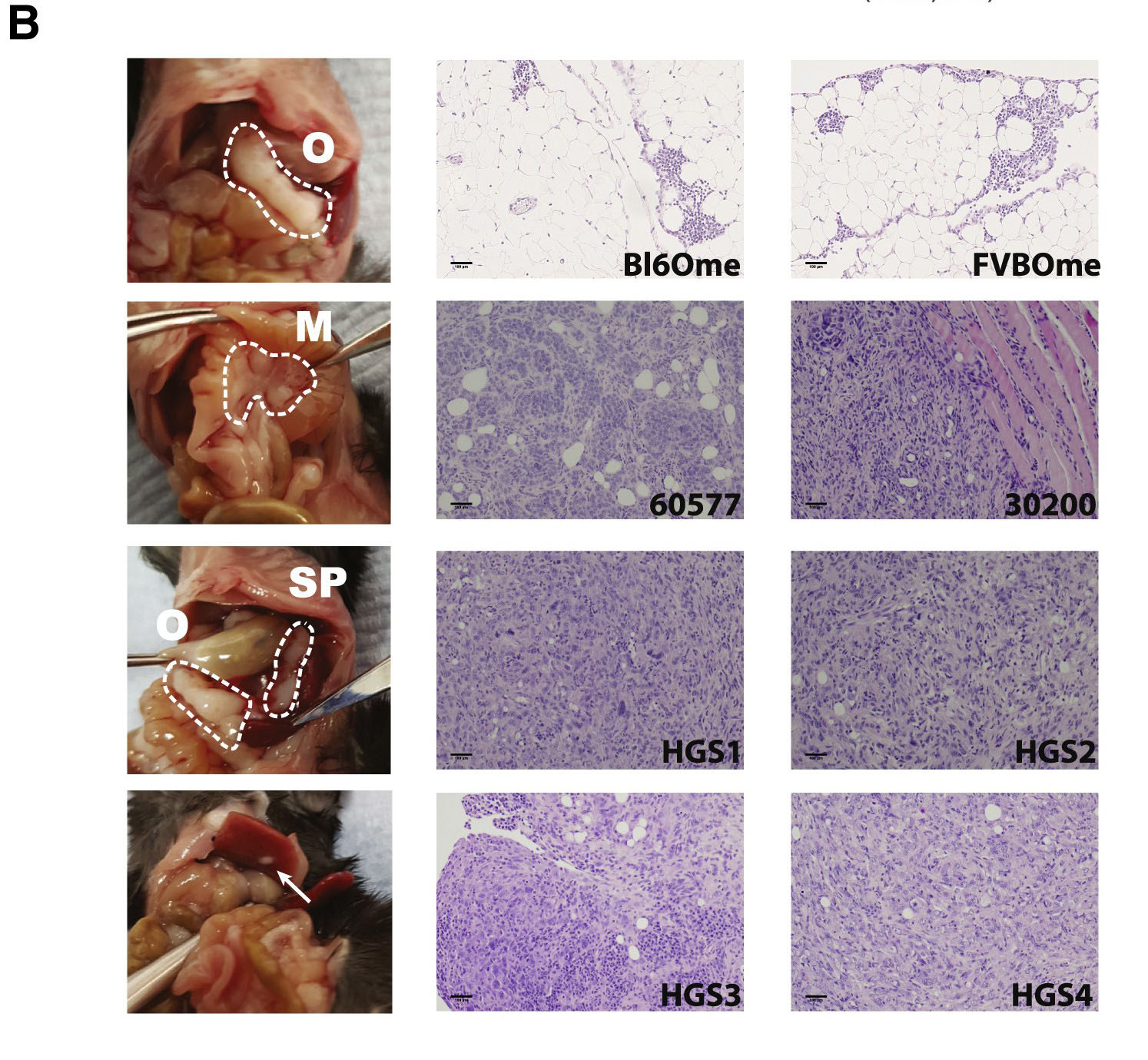
Melanocytes are the cells in mammals that produce pigment (melanin), colouring the hair, skin and irises. They develop from unpigmented precursors, melanoblasts, and are located in the bottom layer of the skin’s epidermis.
The cell lines deposited with CancerTools.org are immortal melanocyte, melanoblast, and neural-crest stem cell lines derived from embryonic mouse skin. These mutant cell lines are used to study the actions of mutated genes, which affect many body systems besides pigment cells. To date, these cell lines have been used in research on topics including cell differentiation, organelle biosynthesis and transport, protein transport, growth control, cancer and many others.
These cell lines therefore not only add value to many areas of pigment cell research including cell biology, developmental biology, molecular biology, genetics, microscopy, physiology, pathophysiology, ageing and cancer, but also to research involving most major organ systems – eyes, ears, and blood, nervous, respiratory, digestive, excretory and skeletal systems, and disorders such as inflammation, thrombosis and allergy among others.
Colour mutations in mice often have an orthologous mutation in humans with associated pathological effects. There is ready interchange between the advances in pigmentary genetics in the mouse and human, which increases the relevance of these cell lines. Thus, a very broad range of body systems, cellular mechanisms and disorders is addressed by this collection of cell lines.
The majority of cell lines with pigmentary mutations were derived from the C57BL/6J strain mice to exclude confounding differences due to strain background. This is a benefit over human cell lines that have many polymorphisms that can affect biological processes independent of known mutations. Several melanocyte (melan-Ink4a-Arf) lines on the C57BL/6J strain background (genotype a/a) were deposited with CancerTools.org. These and some other deposited lines have mutations at the Ink4a-Arf locus that make spontaneous immortalisation routine. Other lines were derived by rare spontaneous immortalisation. Melan-Ink4a-Arf lines are used in applications like the widely used cell line melan-a that immortalised spontaneously. When mutant cell lines were established from mice of other backgrounds, the corresponding wild-type cell lines were established from littermate controls.
Discover more about Dr. Elena Sviderskaya's cell lines:
1https://www.sgul.ac.uk/profiles/elena-sviderskaya#overview