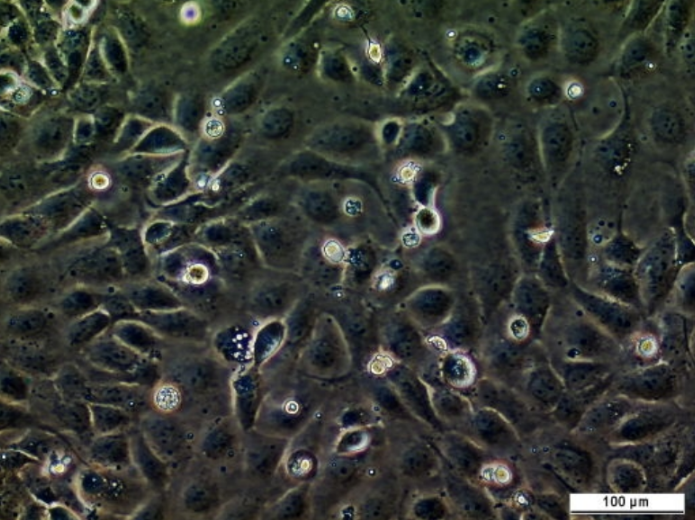
Olaparib-resistant PEO1-OR cell line
Contributors
| Inventor | Institute |
|---|---|
| Aneta Rogalska, Łukasz Biegała, Agnieszka Marczak | University of Lodz |
| Cat. #: | 162283 |
|---|---|
| Cancer types: | Ovarian cancer |
| Organism: | Human |
| Tissue: | Ovary |
| Gender: | Female |
| Morphology: | Epithelial |
| Growth properties: | Adherent |
| Cancer Types In Detail: | Gynaecologic cancer;High Grade Ovarian;High-Grade Serous Ovarian (HGSOC) |
| Primary citation: | Biegala et al. (2023) Cells, 12(7), 1038, PMID: 37048111 |
| Alternate name: | PEO1-OR; PEO1-OR cell line; PEO1-OR cells; olaparib-resistant PEO1-OR ovarian cancer cell line |
|---|---|
| Product description: | The PEO1-OR cell line is a human ovarian cancer cell line with acquired resisatnt to olaparib established from the parental PEO1 high-grade serous ovarian cancer cell line through continuos exposure to step-wise increasing doses of olaparib. PEO1-OR cell line harbours BRCA2 secondary mutation (c.4964A > T) that reverse truncating BRCA2 mutation originating from the parental PEO1 cell line (c.4965C > G; p.Y1655*) and expresses full-length BRCA2 variant (c.[4964A > T; 4965C > G], p.Y1655L). PEO1-OR cells are also less sensitive to ceralasertib (ATR inhibitor) and MK-8776 (CHK1 inhibitor) compared to parental PEO1 cells. PEO1-OR cell line is a powerful in vitro model for studying the molecular and cellular mechanisms of acquiring and overcoming resistance to olaparib in high-grade serous ovarian cancer. This model also allows testing alternate therapies for olaparib-resistant ovarian cancer. |
| Gender: | Female |
| CRISPR: | No |
| Production details: | PEO1-OR cell line is an olaparib-resistant subline of PEO1 cell line developed by addition of a stepwise increase in olaparib to the growth medium of the parental line. |
| Additional notes: | PEO1-OR cells are grown and subcultured continually in the absence of olaparib. Please be aware that the originating laboratory of the PEO1 cell line acknowledges that PEO1 is both genetically unstable and derived from a heterogeneous population that was already present in the patient at the time of biopsy. This is evident in the literature (Cooke et al., 2010). Genetic differences within the PEO1, PEO4 and PEO6 cell lines suggest that PEO4 and PEO6 are not direct descendants of PEO1 but have diverged from a common ancestor. |
| Parental cell line: | PEO1 |
| Model description: | Key mutations determined by whole-exome sequencing using NGS: TP53 (c.731G A; p.G244D); BRCA2 (c.[4964A>T; 4965C>G], p.Y1655L)) |
| Disease: | Cancer |
| Initial handling information: | Thaw cells rapidly in a 37°C water bath for up to 2 minutes, until a small amount of ice remains. Slowly add 1 mL of pre-warmed growth medium (37°C) to the thawed cells. Gently reconstitute the cell pellet by pipetting, then transfer the resuspended cells to a centrifuge tube and add 10 mL of pre-warmed growth medium (37°C). Pellet the cells by centrifugation (300×g, 5 min, RT) to remove the storage medium containing DMSO. After the centrifugation, cell pellet should be visible. Aspirate the supernatant, add 1 mL of growth medium, and gently resuspend the cells. Transfer the cells to into the appropriate culture vessel (e.g. T75 flask) containing medium that has been equilibrated in the incubator for a few minutes (37°C, 5% CO2). Optionally, take a small aliquot to assess cell viability. Rock the flask to distribute the cells evenly and place it in the incubator (37°C, 5% CO2). Thawed cells should be plated at high density to optimize recovery. Examine the cells microscopically after 24 hours and sub-culture as necessary. |
|---|---|
| Format: | Frozen |
| Shipping conditions: | Growth medium (RPMI 1640 culture medium + 2 mM glutamine + 25 mM HEPES + 10% heat inactivated FBS) + 10% sterile DMSO |
| Growth medium: | RPMI 1640 culture medium with 2 mM glutamine and 25 mM HEPES supplemented with 10% heat inactivated FBS |
| Subculture routine: | Split sub-confluent cultures (70–90%) every 2 to 4 days using a 0.25% trypsin-EDTA solution. Briefly, remove growth medium, wash cells with PBS without Ca2+/Mg2+, add trypsin-EDTA solution (approximately 1.5–2 mL per 75 cm2 of surface area), place flask in the incubator and leave it for 5–10 minutes (37°C, 5% CO2). Once the cells are detached, add fresh growth medium, and transfer the required number of cells to a new flask. Optionally, after trypsin inactivation, centrifuge cells (300×g, 5 min, RT), resuspend cell pellet in growth medium and passage required number of cells to a new flask. Recommended culture vessel: cell culture treated flasks with filter caps. As a general guide, 70–90% confluent cells should be split in approximately 1:4, 1:6 or 1:12 ratios to reach appropriate confluency after 2, 3 or 4 days. However, splitting ratio may vary depending on cell line handling and culture conditions. The complete formulation of the recommended trypsin-EDTA solution is: KCl (75 mg/L), KH2PO4 (60 mg/L), NaHCO3 (350 mg/L), NaCl (8000 mg/L), Na2HPO4 · 7H2O (90 mg/L), D-glucose (1000 mg/L), EDTA–4Na · 2H2O (380 mg/L), phenol red (10 mg/L), trypsin (2500 mg/L), pH 7.2 to 8. |
| Temperature: | 5% CO2 |
| Atmosphere: | Dry ice |
| Storage medium: | Liquid nitrogen, 2×10^6 cells/vial |
| Mycoplasma free: | Yes |
| Biosafety level: | 2 |
| References: |
Biegala et al. (2023) Cells, 12(7), 1038, PMID: 37048111 |
|---|
Images
View GalleryRelated Tools for Cell lines
| Cat. # | Tool Name | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 151014 | MCF7 AREc32 Cell Line |
Key Info
MCF7 AREc32 Cell Line
|
View Tool | |||||||||||||
| 151463 | Omega-E Packaging Cell Line |
Key Info
Omega-E Packaging Cell Line
|
View Tool | |||||||||||||
| 151629 | MEF Raf1 FF/FF KI Cell Line |
Key Info
MEF Raf1 FF/FF KI Cell Line
|
View Tool | |||||||||||||
| 151448 | CMT 167 Cell Line |
Key Info
CMT 167 Cell Line
|
View Tool | |||||||||||||
| 151576 | MEF RAF1 KO Cell Line |
Key Info
MEF RAF1 KO Cell Line
|
View Tool | |||||||||||||