Anti-HPV16E2 [HPV16E2]
Cervical intraepithelial neoplasia (CIN) is caused by human papillomavirus (HPV) infection and is the precursor to cervical carcinoma. The completion of the HPV productive life cycle depends on the expression of viral proteins. Initiation of the viral productive replication requires expression of the E2 viral protein that cooperates with the E1 viral DNA helicase. A […]
Contributors
| Institute |
|---|
| A*STAR Accelerate Technologies Pte Ltd |
| Cat. #: | 153202 |
|---|---|
| Tool sub type: | Primary antibody |
| Unit size: | 100 ug |
| Cancer types: | Cervical cancer |
| Research Fields: | Cancer;Cell biology;Microbiology |
| Application: | IHC ; WB |
| Target: | HPV16E2 |
| Reactivity: | Human |
| Clone: | HPV16E2 |
| Host: | Rabbit |
| Class: | Polyclonal |
| Alternate name: | Human papillomavirus type 16; Regulatory protein E2 |
|---|---|
| Product description: | Cervical intraepithelial neoplasia (CIN) is caused by human papillomavirus (HPV) infection and is the precursor to cervical carcinoma. The completion of the HPV productive life cycle depends on the expression of viral proteins. Initiation of the viral productive replication requires expression of the E2 viral protein that cooperates with the E1 viral DNA helicase. A novel and important role for the HPV16-E2 protein in controlling host cell cycle during malignant transformation was recently revealed. Xells expressing HPV16-E2 in vitro were arrested in prophase alongside activation of a sustained DDR signal. In vivo, HPV16-E2 protein is present in cells that express both mitotic and DDR signals specifically in CIN3 lesions, immediate precursors of cancer, suggesting that E2 may be one of the drivers of genomic instability and carcinogenesis in vivo. |
| Conjugation: | Unconjugated |
| Immunogen: | GST-HPV16 E2 (209365) |
| Immunogen Uniprot ID: | TBC |
| Target background: | Cervical intraepithelial neoplasia (CIN) is caused by human papillomavirus (HPV) infection and is the precursor to cervical carcinoma. The completion of the HPV productive life cycle depends on the expression of viral proteins. Initiation of the viral productive replication requires expression of the E2 viral protein that cooperates with the E1 viral DNA helicase. A novel and important role for the HPV16-E2 protein in controlling host cell cycle during malignant transformation was recently revealed. Xells expressing HPV16-E2 in vitro were arrested in prophase alongside activation of a sustained DDR signal. In vivo, HPV16-E2 protein is present in cells that express both mitotic and DDR signals specifically in CIN3 lesions, immediate precursors of cancer, suggesting that E2 may be one of the drivers of genomic instability and carcinogenesis in vivo. |
|---|
| Format: | Liquid |
|---|---|
| Concentration: | 0.9-1.1 mg/ml |
| Storage buffer: | Whole serum |
| Storage conditions: | -15° C to -25° C |
| Shipping conditions: | Dry ice |
| References: |
Xue et al. 2015. Oncotarget. 6(33):34979-91. PMID: 26474276. HPV16-E2 induces prophase arrest and activates the cellular DNA damage response in vitro and in precursor lesions of cervical carcinoma. Xue et al. 2010. Cancer Res. 70(13):5316-25. PMID: 20530671. HPV16 E2 is an immediate early marker of viral infection, preceding E7 expression in precursor structures of cervical carcinoma. |
|---|
Images
View GalleryProduct Images
b1ec6bdf-f092-4ac2-945d-eac28efbc78f.jpg
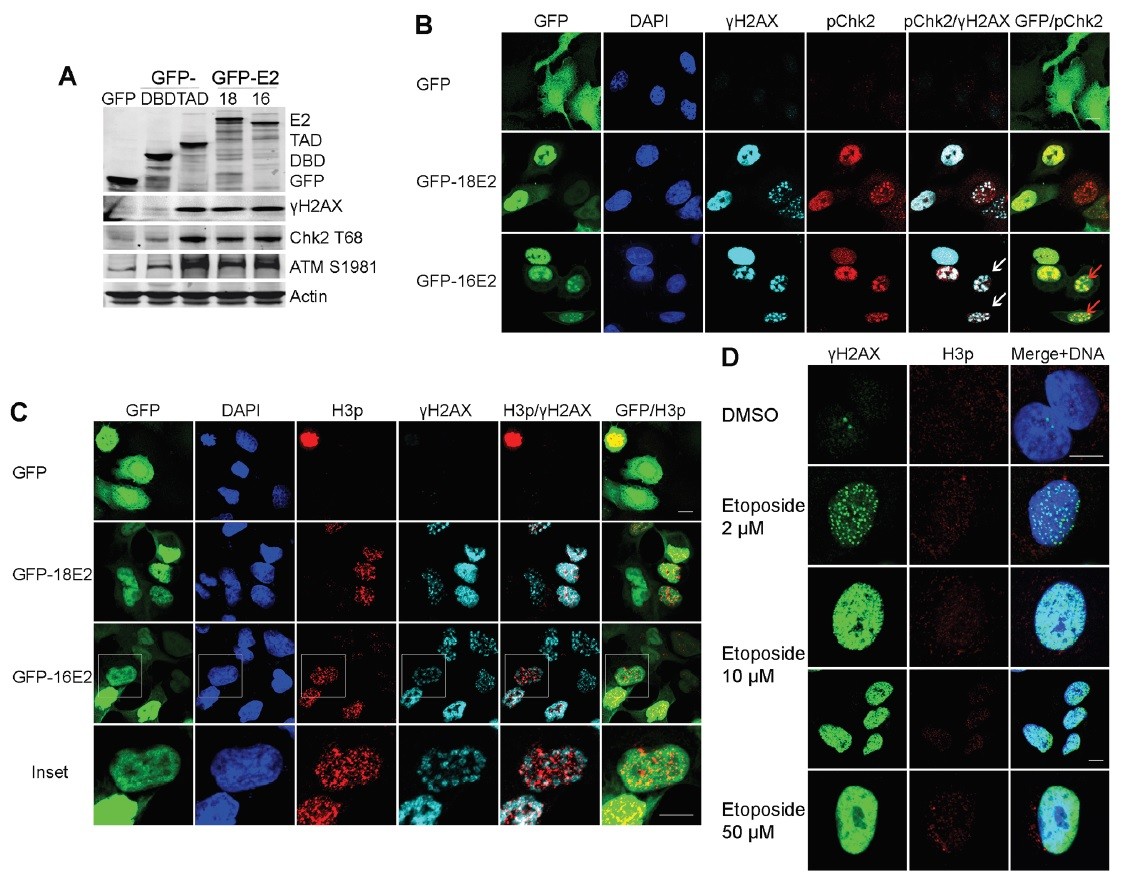
Figure. E2 induces a DNA damage signal in A549 cells arrested in prophase. A. Western blot analyses of lysates from A549 cells expressing GFP-16E2; GFP-18E2; HPV18E2 transactivation domain (GFP-18E2TAD) or the DNA binding domain (GFP-18E2DBD); probed with antibodies against GFP; ?H2AX; Chk2T68 and ATMS1981. B. Confocal microscopy of E2-expressing A549 cells double labeled with anti-?H2AX (light blue) and anti-Chk2T68 (red). White arrows

Related Tools for Antibodies
| Cat. # | Tool Name | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 151016 | Anti-CyclinA [E72.1] |
Key Info
Anti-CyclinA [E72.1]
|
View Tool | |||||||||||||||||||
| 151019 | Anti-Mos [S3.1] |
Key Info
Anti-Mos [S3.1]
|
View Tool | |||||||||||||||||||
| 151024 | Anti-Hairy [1/24] |
Key Info
Anti-Hairy [1/24]
|
View Tool | |||||||||||||||||||
| 151031 | Anti-RuvA [RuvA 12C6] |
Key Info
Anti-RuvA [RuvA 12C6]
|
View Tool | |||||||||||||||||||
| 151028 | Anti-Cytochrome P450 2C2, 2B1/2 [h7] |
Key Info
Anti-Cytochrome P450 2C2, 2B1/2 [h7]
|
View Tool | |||||||||||||||||||
