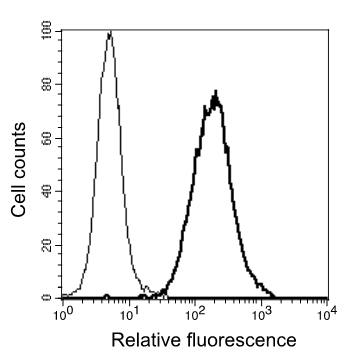
Anti-CEACAM21 [1E4]
Contributors
| Inventor | Institute |
|---|---|
| Bernhard B. Singer | LeukoCom |
| Cat. #: | 153999 |
|---|---|
| Tool sub type: | Primary antibody |
| Unit size: | 100 ug |
| Research Fields: | Cancer;Cell biology |
| Application: | ELISA ; FACS ; WB |
| Target: | Human CEACAM 21 |
| Reactivity: | Human |
| Host: | Mouse |
| Class: | Monoclonal |
| Conjugation: | Unconjugated |
|---|---|
| Isotype: | IgG1 kappa |
| Immunogen: | Recombinant soluble human CEACAM 21-Fc produced in HEK293 Transfectants |
| Myeloma used: | NS1/0 |
| Format: | Liquid |
|---|---|
| Concentration: | 0.9-1.1 mg/ml |
| Storage buffer: | PBS with 0.02% azide |
| Storage conditions: | -15° C to -25° C |
| Shipping conditions: | Dry ice |
| References: |
Tchoupa et al. 2014. Cell Commun Signal. 12:27. PMID: 24735478. Signaling by epithelial members of the CEACAM family – mucosal docking sites for pathogenic bacteria. Genetics of sputum gene expression in chronic obstructive pulmonary disease. Qiu et al. 2011. PLoS One. 6(9):e24395. PMID: 21949713. Zebhauser et al. 2005. Genomics. 86(5):566-80. PMID: 16139472. Identification of a novel group of evolutionarily conserved members within the rapidly diverging murine Cea family. Fukuda, I., Yamakado, M. & Kiyose, H. Journal of Medical Systems (1998) 22: 89. |
|---|
Images
View GalleryRelated Tools for Antibodies
| Cat. # | Tool Name | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 151015 | Anti-CyclinA [E70.1] |
Key Info
Anti-CyclinA [E70.1]
|
View Tool | |||||||||||||||||||
| 151036 | Anti-ICAM1 [15.2] |
Key Info
Anti-ICAM1 [15.2]
|
View Tool | |||||||||||||||||||
| 151020 | Anti-CyclinB1 [V143.1] |
Key Info
Anti-CyclinB1 [V143.1]
|
View Tool | |||||||||||||||||||
| 151028 | Anti-Cytochrome P450 2C2, 2B1/2 [h7] |
Key Info
Anti-Cytochrome P450 2C2, 2B1/2 [h7]
|
View Tool | |||||||||||||||||||
| 151035 | Anti-CD8 [14] mAb |
Key Info
Anti-CD8 [14] mAb
|
View Tool | |||||||||||||||||||