Non-small cell lung cancer (NSCLC) is not a static disease — it is an evolving ecosystem shaped by clonal diversity and therapeutic pressure. In response to this challenge, Dr Robert Hynds and Dr David Pearce from University College London (UCL) have developed a unique collection of evolution-informed PDX models within the Lung TRACERx programme. These models capture tumour heterogeneity with exceptional clarity. In this blog, we explore how these models were generated, what makes them distinct, and why they offer scientists a powerful platform for studying tumour evolution and validating new therapies in vivo.

Rethinking lung cancer models to address tumour heterogeneity
Lung cancer remains the leading cause of cancer-related death worldwide, accounting for nearly 1.8 million deaths each year (1). Many of these cases are NSCLC, a disease that has proven difficult to treat despite advances in therapy. One key reason is intratumour heterogeneity (IH) – the hidden diversity within each tumour.
Rather than being a uniform disease, NSCLC is a dynamic ecosystem composed of multiple genetically and phenotypically distinct cell populations that adapt under therapeutic pressure. This diversity is what drives treatment resistance, contributes to sampling bias from single biopsies, and ultimately limits the long-term success of targeted therapies.
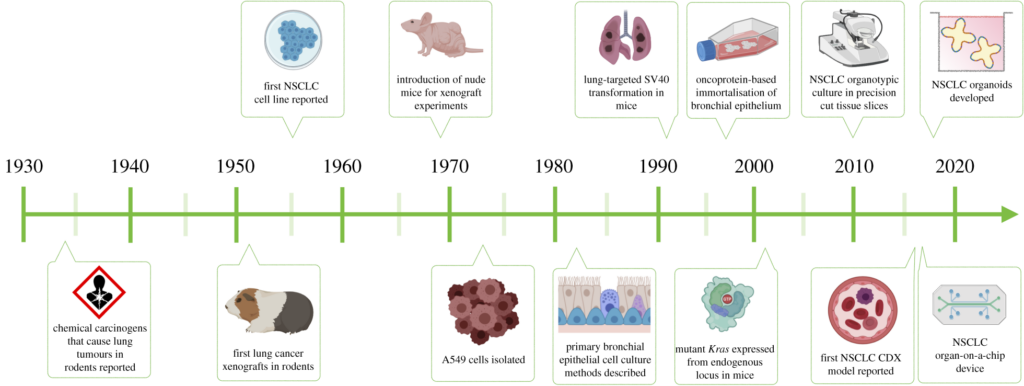
Our understanding of NSCLC has been improved by various tumour models, including immortalised cell lines, primary cell cultures, conventional animal models and more (Fig. 1). However, these often fall short in recapitulating patient tumour architecture, microenvironment, or clonal diversity. As a result, there is a growing need to rethink how we model NSCLC — developing systems that better capture tumour complexity and heterogeneity.
“Capturing intratumour heterogeneity is critical because NSCLC evolves and evades therapy through the selection and expansion of diverse subclonal populations. Models that fail to represent this diversity provide only a partial view of the disease and therapy responses.”
Dr. David Pearce, UCL
Redefining preclinical cancer models with PDXs
To overcome limitations, the field is moving towards more clinically representative, patient-centric 3D models, such as patient-derived xenografts (PDXs). Developed by implanting patient tumour tissue into immunocompromised mice, PDXs retain key biological features of the original tumour, including aspects of the tumour microenvironment (TME) and IH.
Unlike traditional 2D cell lines, which often represent highly selected subclones adapted to specific culture conditions, PDXs preserve the heterogeneous cell populations present in patient tumours. This makes them particularly valuable for studying NSCLC, where subclonal diversity drives tumour progression and therapeutic resistance.
Early-passage PDXs can also maintain elements of patient-derived stroma, influencing tumour growth and treatment response, and often show drug response patterns that better reflect clinical outcomes. Despite limitations — including stromal replacement over time, cost, and technical complexity — PDXs remain one of the most powerful in vivo platforms for studying tumour evolution and improving translational research.
TRACERx: mapping the evolution of NSCLC
Lung TRACERx (TRAcking Cancer Evolution through therapy (Rx)) is a landmark study funded by Cancer Research UK and led by Prof. Charles Swanton at UCL and The Francis Crick Institute (3). Taking place over nine years and involving 800 patients across 20 hospitals in the UK, TRACERx represents one of the most comprehensive efforts to understand how NSCLC evolves over time. The study brings together multidisciplinary expertise to integrate clinical, histopathological and genomic data at an unprecedented scale.
Using multi-region longitudinal tumour sampling and genetic analysis by whole-exome sequencing (WES), TRACERx maps how tumours adapt under therapeutic pressure and how IH contributes to disease recurrence and immune evasion. By generating a longitudinal view from diagnosis through treatment and relapse, the study reframes NSCLC as an evolving disease rather than a static snapshot.
This evolutionary perspective will have profound implications on clinical outcomes, such as informing patient stratification and the development of more durable targeted and immune-based therapies.
What Hynds and Pearce built to reflect NSCLC evolution
A total of 48 PDX models were developed from 22 patients in the Lung TRACERx study by Dr Robert Hynds and Dr David Pearce at the UCL Cancer Institute (4). Their work was driven by a clear scientific question: how faithfully do PDX models capture the true genomic complexity of heterogeneous NSCLC tumours?
“Studying viable tumour cells directly from patients remains a major challenge, even with advances in cell line and organoid culture systems. We aimed to develop models that could maintain patient-derived tumour tissue from a wider range of cases and to investigate how well they preserved tumour diversity.”
Dr. David Pearce, UCL
While PDXs are widely used, uncertainties remained about their ability to represent tumour subclonal diversity, withstand genetic drift during engraftment, and avoid sampling bias from single tumour regions (4,5). To address this, Hynds, Pearce and Dr. Ariana Huebner from The Francis Crick Institute paired WES data from multiple regions of TRACERx patient tumours with their matched PDX models. This allowed them to quantify how engraftment shapes clonal representation, determine whether different tumour regions generate distinct PDXs, and measure evolutionary changes occurring in vivo (Figure 2).
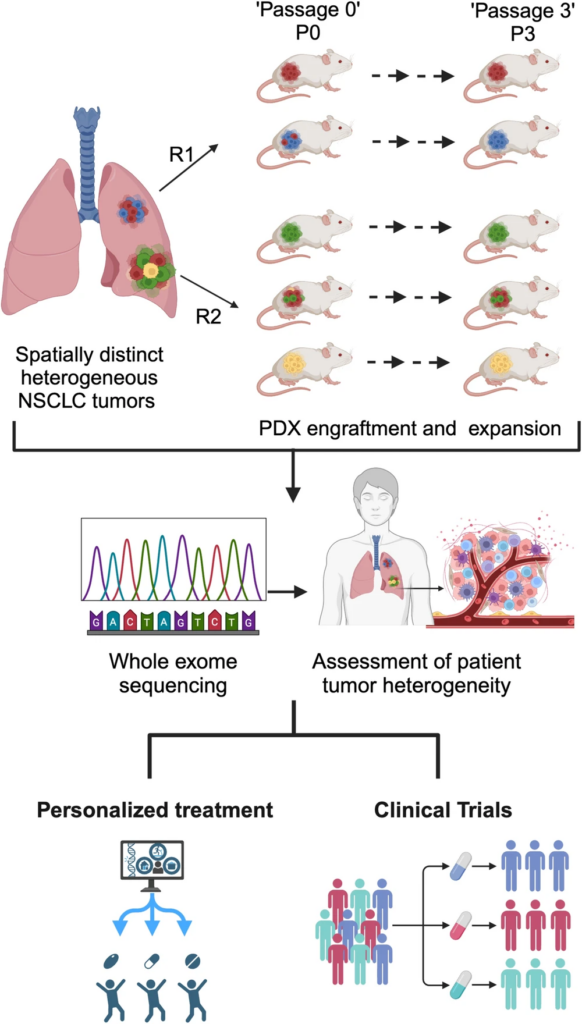
Embedding these PDXs within the TRACERx framework transforms them from standalone xenografts into evolution-informed research models. Each is linked to multi-region sequencing data and long-term clinical follow-up from the original patient tumour, allowing it to be placed within the tumour’s evolutionary history rather than being studied as an isolated sample.
Importantly, generating PDXs from multiple regions of the same tumour enables direct assessment of IH, reveals clonal bottlenecks during engraftment, and allows comparison between models derived from a single patient.
The result is a deeply characterised collection of PDX models that provides a more precise platform for studying clonal selection, resistance mechanisms, and therapeutic response in NSCLC. By combining technical rigour with evolutionary context, Hynds, Pearce and colleagues have established a resource that advances not only model fidelity, but our ability to interpret and translate preclinical findings with confidence.
From tumour evolution to translational insight
The TRACERx NSCLC PDX models were designed to do more than generate transplantable tumours — they were built to capture IH in a structured and traceable way.
To achieve this, spatially distinct regions of the same primary patient tumour were independently sequenced and used to generate matched PDX models. Because different tumour regions often harbour distinct subclones, this multi-region approach allows each PDX to be mapped back to a defined branch of the tumour’s evolutionary tree. In practice, individual models frequently represent a dominant subclone, while multi-region-matched panels from the same patient reflect broader tumour diversity (4).
For researchers, this has direct practical value. If your work focuses on IH, these models enable the study of defined tumour subpopulations within a living in vivo system. You can examine how specific evolutionary branches respond to selective pressure, how resistance emerges from clones, and how clonal dynamics influence treatment outcome.
For drug discovery and translational teams, these PDXs offer a powerful platform for in vivo drug validation. Because each model is rooted in detailed genomic data, you can test novel compounds against tumours with known clonal architecture – allowing for more informed efficacy assessment across distinct subpopulations and rational combination strategies aimed at preventing or delaying treatment resistance.
As a whole, the TRACERx NSCLC PDX collection provide scientists with a toolkit for a clearer path toward therapies designed to outpace tumour evolution.
Making evolution-informed PDX models accessible
Through a continued partnership with UCLB (the commercialisation arm of UCL), 44 of these pioneering TRACERx NSCLC PDX models are now accessible to the global cancer research community through CancerTools.
This collaboration reflects a shared mission: to ensure that high-value research tools do not remain siloed but are made available to fellow researchers who can build on them. At CancerTools, our role is to accelerate cancer discovery and innovation by enabling global access to tools that have already been developed, so researchers do not need to rebuild from scratch. By doing so, we help save time, reduce duplication, and minimise the unnecessary use of funding and precious resources.
“Our collaboration with CancerTools allows us to make these innovative TRACERx NSCLC PDX models accessible to support other lung cancer researchers worldwide. This is an exciting opportunity for research efforts at UCL to have impact in the wider lung cancer research community and to help the development of effective therapies.”
Dr. Rob Hynds, UCL
Through this partnership, the TRACERx team provide fellow scientists with reliable models to interrogate tumour heterogeneity, test therapeutic hypotheses and explore resistance mechanisms with greater confidence.
The road ahead in NSCLC research and treatment
Lung TRACERx continues to deepen our understanding of NSCLC evolution. Yet important questions remain. For example, which subclones drive relapse and metastasis? How can resistance trajectories be predicted earlier? The TRACERx PDX models provide a practical platform to help address these questions, enabling functional testing of evolution-informed hypotheses and in vivo validation of therapies within defined clonal contexts.
As modelling approaches continue to evolve, including the development of more immune-competent systems, evolution-informed PDXs remain an important bridge between genomic insight and translational research. By capturing tumour diversity within a structured evolutionary framework, Hynds, Pearce and colleagues have helped drive a broader shift in oncology — where tumour evolution is not just observed, but anticipated, shaping how therapies are designed from the very beginning.
References
- World Health Organization: Lung Cancer. https://www.who.int/news-room/fact-sheets/detail/lung-cancer
- Hynds, R.E., et al., 2021. Open Biology. 11 (1): 200247. PMID: 33435818
- Jamal-Hanjani, M., et al., 2014. PLoS Biology. 12(7), e1001906. PMID: 25003521
- Hynds, R.E., et al., 2024. Nat Communications. 15, 4653. PMID: 38821942
- Jin, J., et al., 2023. Cancers. 15(17):4352. PMID: 37686627
- Sunil, H.S., et al., 2024. Nature Communications. 15, 4652. PMID: 38821926.
