Exclusive preclinical models of high-grade serous ovarian cancer
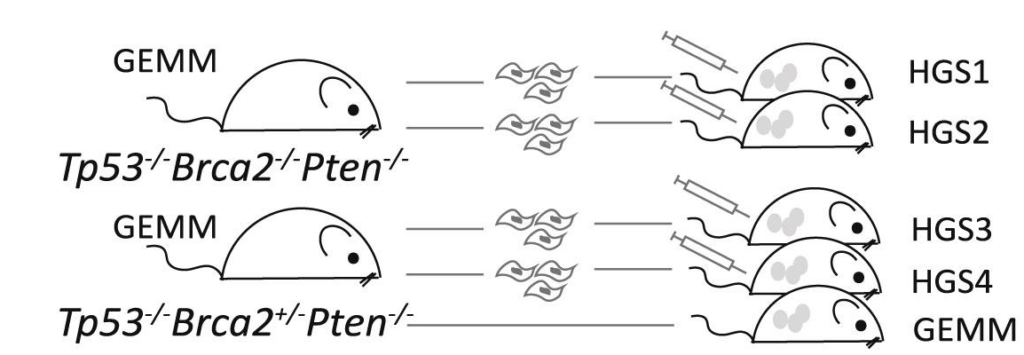
The HGS cell line series (HGS1, HGS2, HGS3 and HGS4) reproduces the genetic drivers, stromal interactions, and immune complexity of human high-grade serous ovarian cancer (HGSOC). Developed in immunocompetent mice, these models enable the study of tumour–microenvironment interactions and provide a translational platform for testing targeted and immunotherapies.
Each line carries distinct genetic alterations and microenvironmental features, together capturing the heterogeneity of HGSOC and its variable treatment responses — a limitation of traditional models.
Meet the cell lines
The HGS 1–4 panel reflects distinct genetic contexts of HGSOC, providing a versatile system to study therapy-sensitive and resistant-like disease.
- HGS1 & HGS2 are Tp53-/-, Pten-/-, Brca2-/- and hence expected to be homologous recombination (HR) deficient and more responsive to platinum and PARP inhibitors, consistent with BRCA2-mutant patient tumours.
- HGS3 & HGS4 are Tp53-/-, Pten-/-, Brca2+/-, and hence partially HR competent, modelling resistant-like biology observed in patient subgroups.
Together, the four models span the clinical spectrum of HGSOC, from therapy-sensitive to resistant-like contexts, enabling preclinical studies that more closely predict patient outcomes.
Core features of HGS1–4 cell lines
- Genetic fidelity: engineered with key alterations observed in human HGSOC, including Tp53, Pten, Brca2 mutations
- Tumour growth and dissemination: form tumours in the omentum of immunocompetent mice, reflecting metastatic spread seen in patients
- Histopathological similarity: reproduce hallmarks of HGSOC such as cellular atypia, papillary/solid growth, and high mitotic activity
- Transcriptomic alignment: gene expression profiles overlap significantly with human tumours, capturing pathways such as epithelial–mesenchymal transition
Data highlights
Data published in Cell Reports (Maniati et al., 2020) show that tumours generated by the HGS1–4 cell lines reproduce key genetic drivers, histopathological architecture, and immune–stromal features of patient HGSOC, including:
Human-like tumour architecture and composition
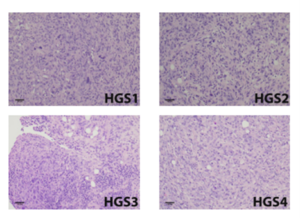
Histological sections from HGS-derived tumours reveal stromal–epithelial organisation and growth patterns (papillary, solid) with high mitotic activity, closely paralleling human HGSOC biopsies, enabling more faithful modelling of tumour–stromal interactions.
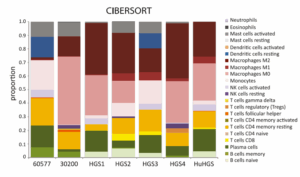
Translationally relevant immune profiles support evaluation of immunotherapy and targeted approaches
Immune profiling (CIBERSORT) shows that HGS tumours mirror the heterogeneity of human HGSOC, with comparable proportions of T-cell, B-cell, and myeloid populations, providing a translationally relevant platform for evaluating immunotherapy and targeted treatment responses.
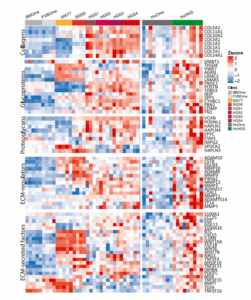
Transcriptional signatures linked to chemotherapy response
RNA-seq profiling of HGS1–4 tumours reveal expression signatures associated with platinum sensitivity and resistance, enabling studies of therapy-responsive and refractory disease
